Studying the microbial symbiosis in its natural context
Species are not independent. They are shaped by the ecological interdependence among members of their communities and the evolutionary history from which they arise. The interplay between Ecology and Evolution gives rise to the biodiversity we observe in nature, with interactions spanning multiple levels of biological organization, from genes to populations to communities.
A central question we ask is to what extent this diversity is driven by
- deterministic processes, such as natural selection and environmental filtering
- stochastic processes, such as historical contingency and genetic drift
Plant–microbe symbioses, together with mobile symbiosis genes, provide a clear example of such interdependent systems. In the legume–rhizobium–MGE (mobile genetic element) system, diversity emerges at each level, including host species and genotypes, rhizobial species and strains, and sequence and structural variation in MGEs.
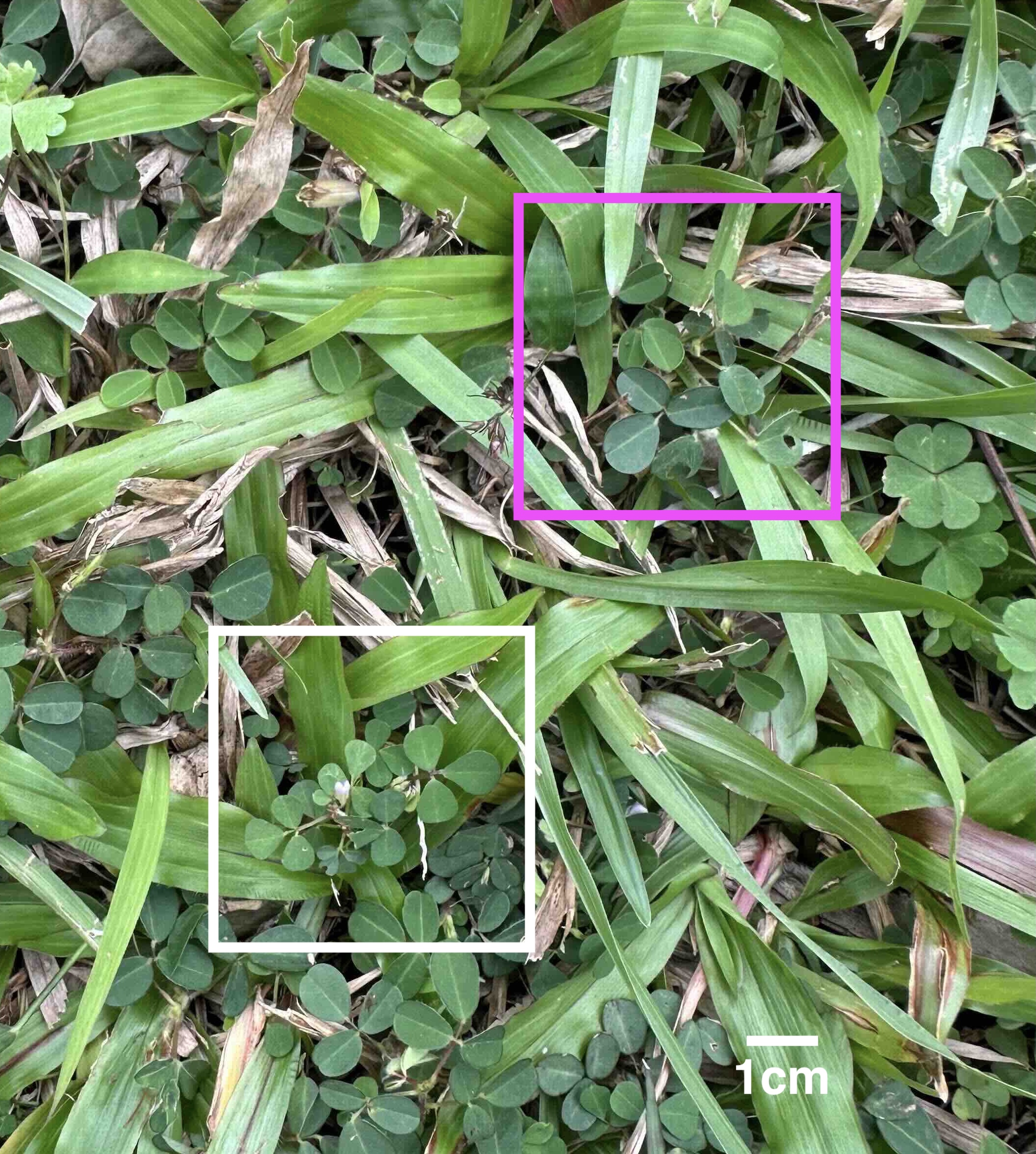
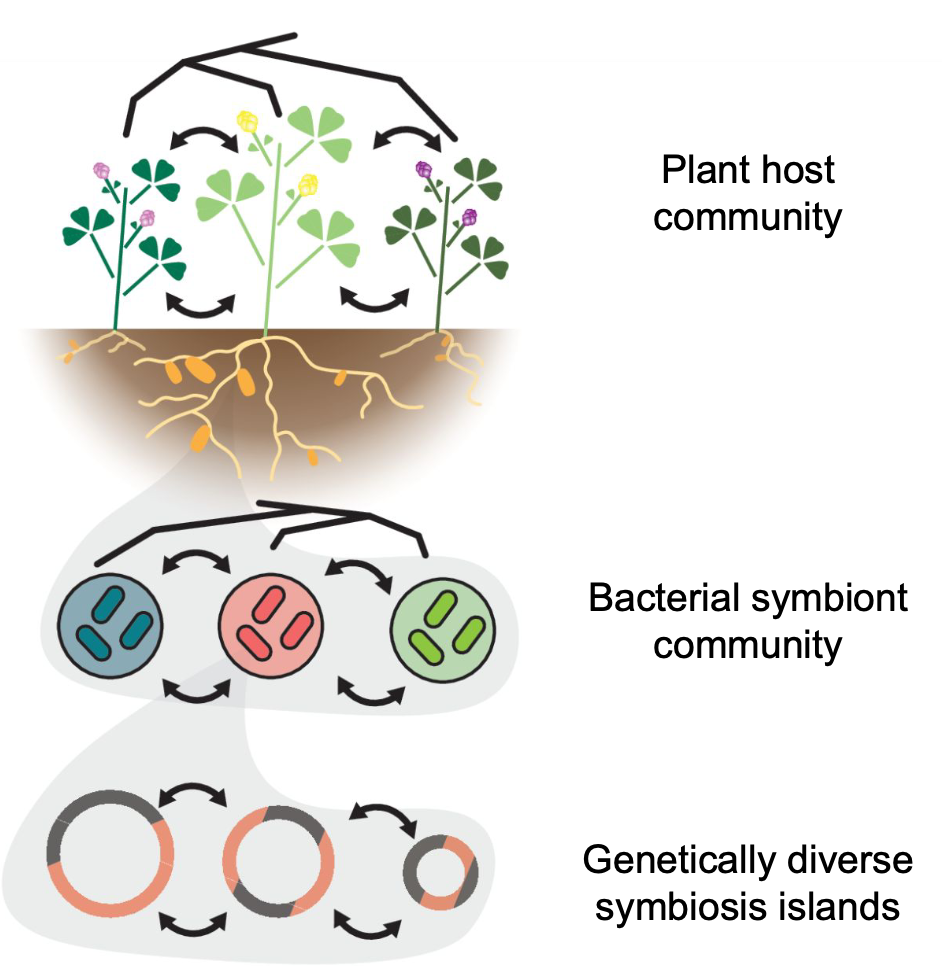

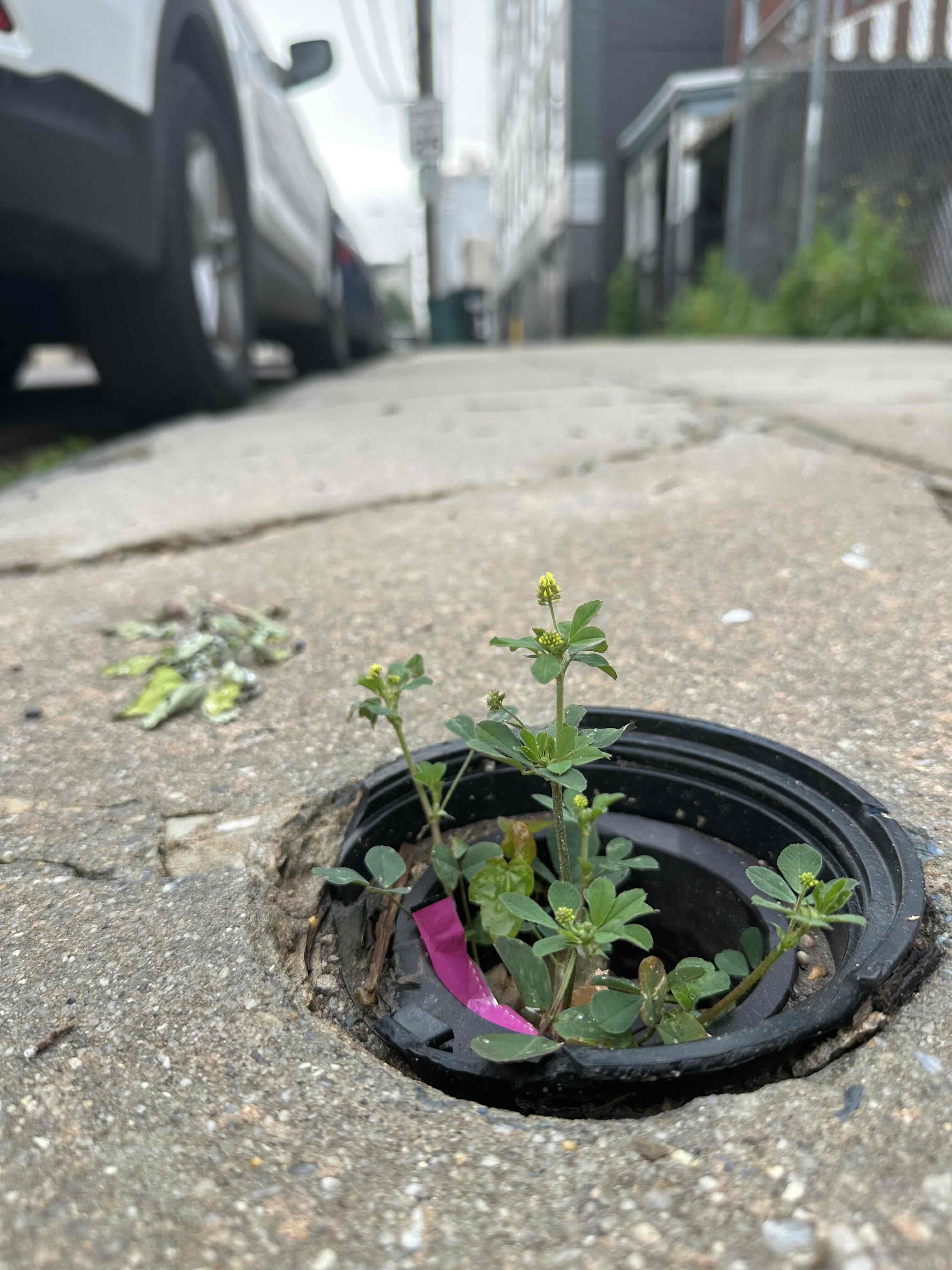
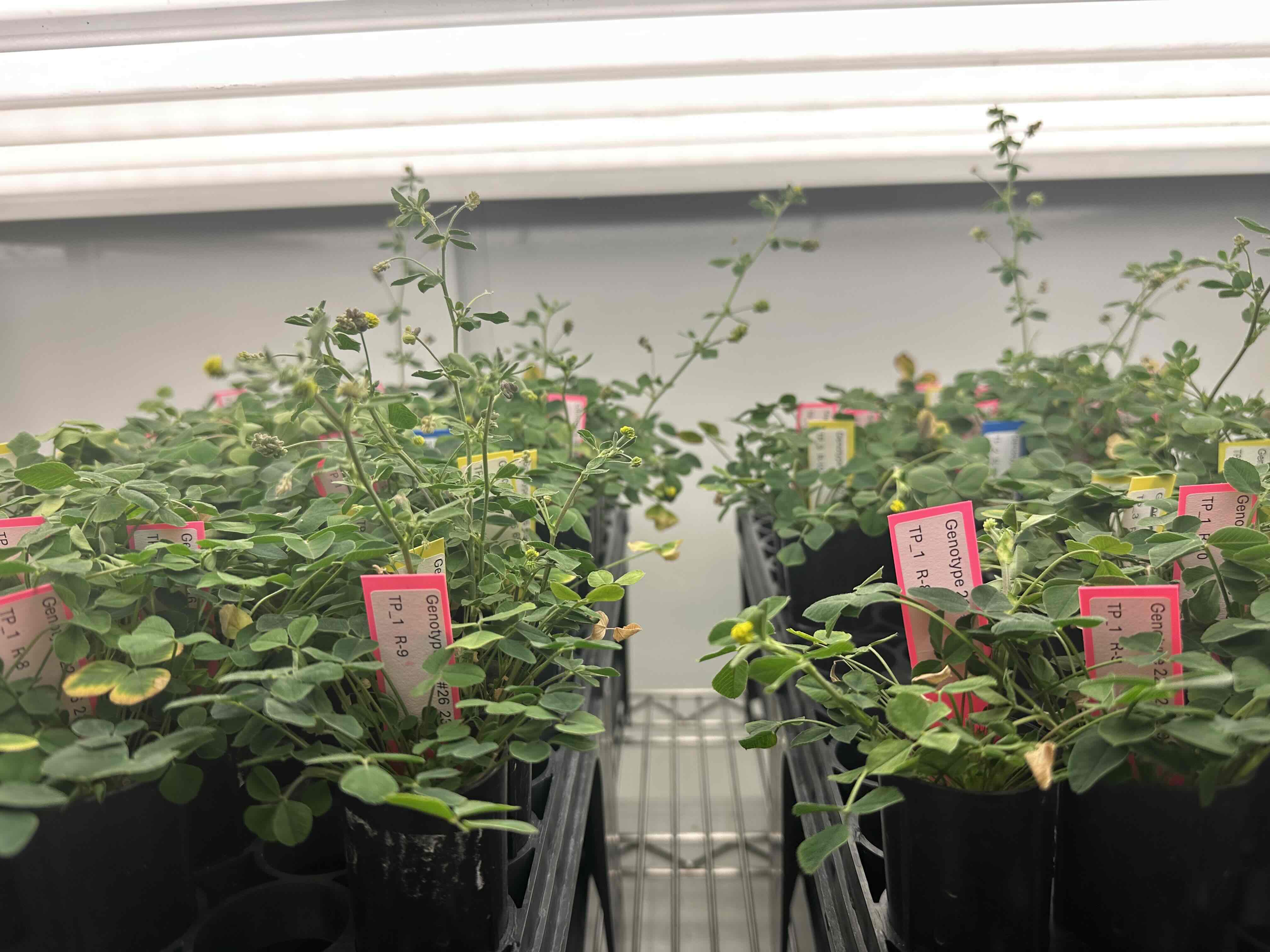
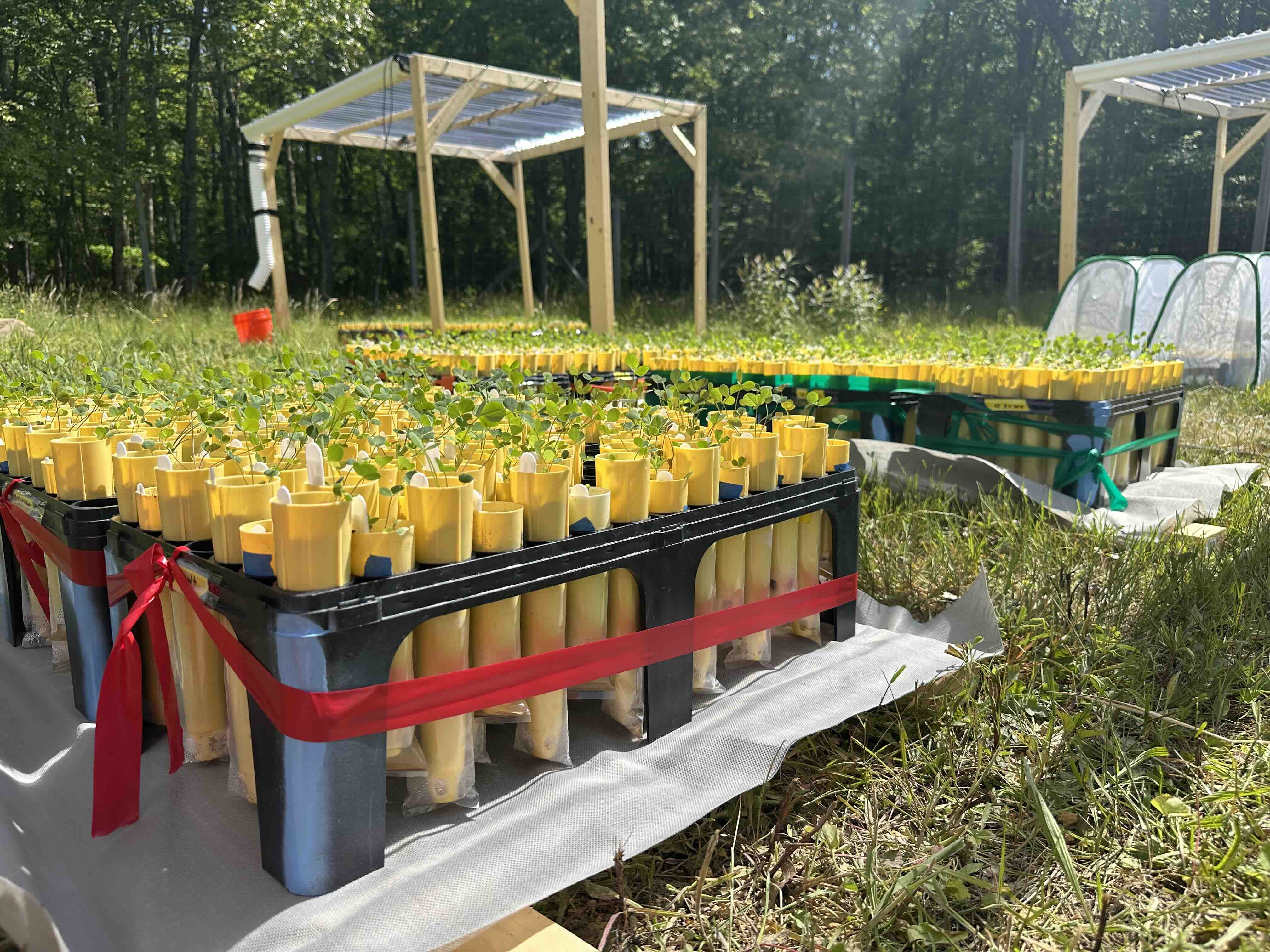
We study a legume system native to Taiwan, the genus Grona 假地豆屬, and its associated rhizobial symbionts.
- Grona species are widely distributed across low-elevation regions of Taiwan
- They are common in urban, agricultural, and semi-natural environments
- They form symbioses with diverse rhizobial taxa, including Bradyrhizobium, Rhizobium, and Sinorhizobium
Together, this system provides a natural framework for studying how symbiosis evolves and functions across heterogeneous environments, including those shaped by human activity.
Evolution of Host–Microbe Symbiosis
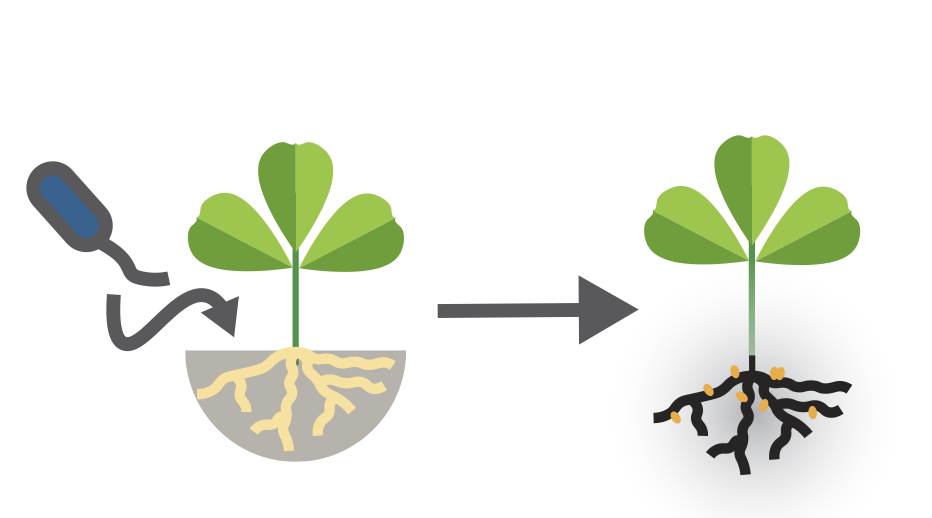
How does the free-living stage shape the evolution of facultative symbionts that regularly associate with hosts?
Adaptation during the free-living stage leaves distinct biogeographic and genomic signatures in facultative symbiotic microbes associated with a widely distributed host.
Population genomics, growth assays, phylogenetic comparative methods.
Facultative symbiosis, mutualism, rhizobia.
Facultative symbionts alternate between host-associated and free-living stages. We study how environmental adaptation outside the host drives genomic divergence and population structure, providing new insights into the evolution of microbial symbiosis.
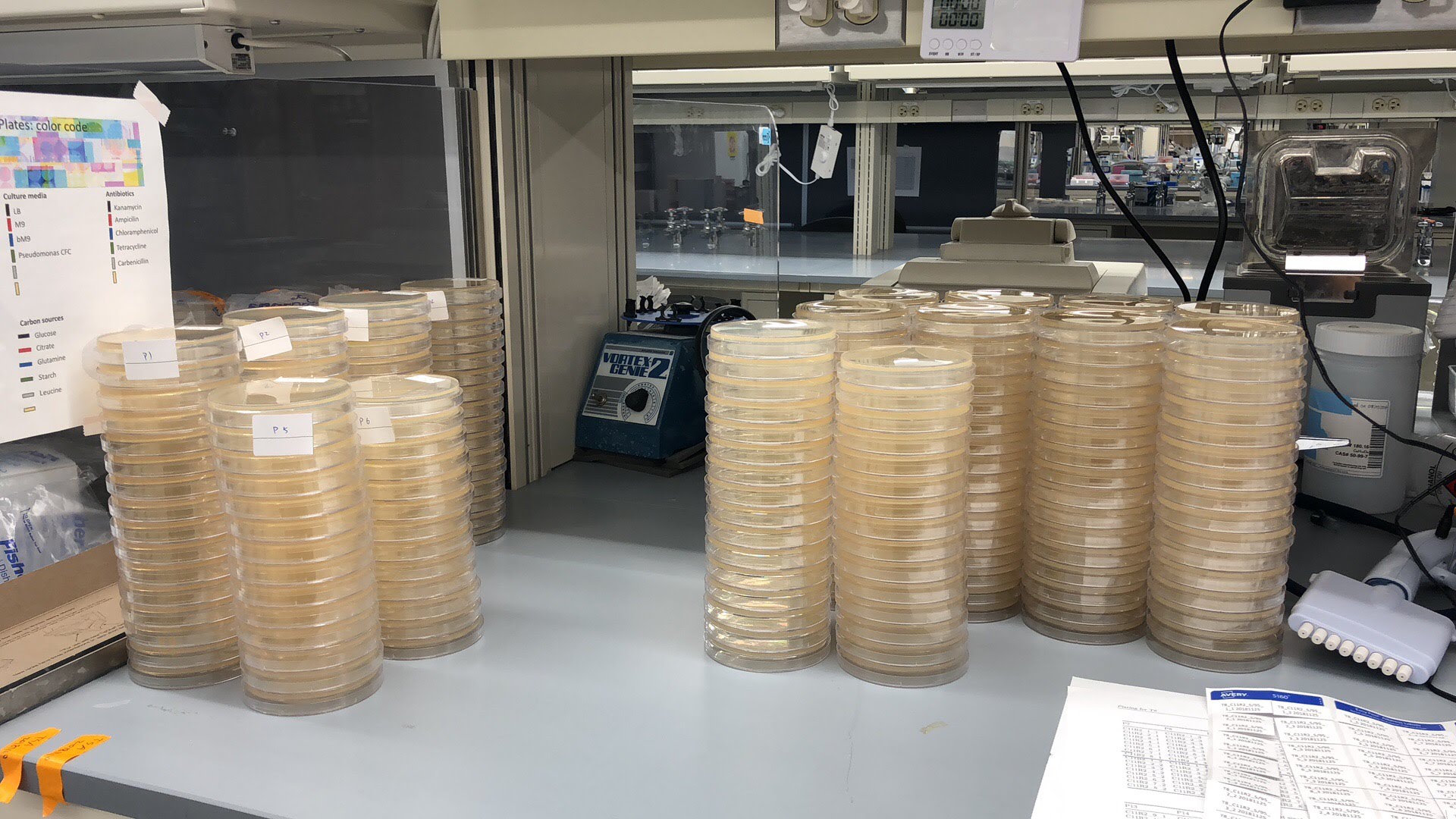
Emergent Coexistence & Community Assembly
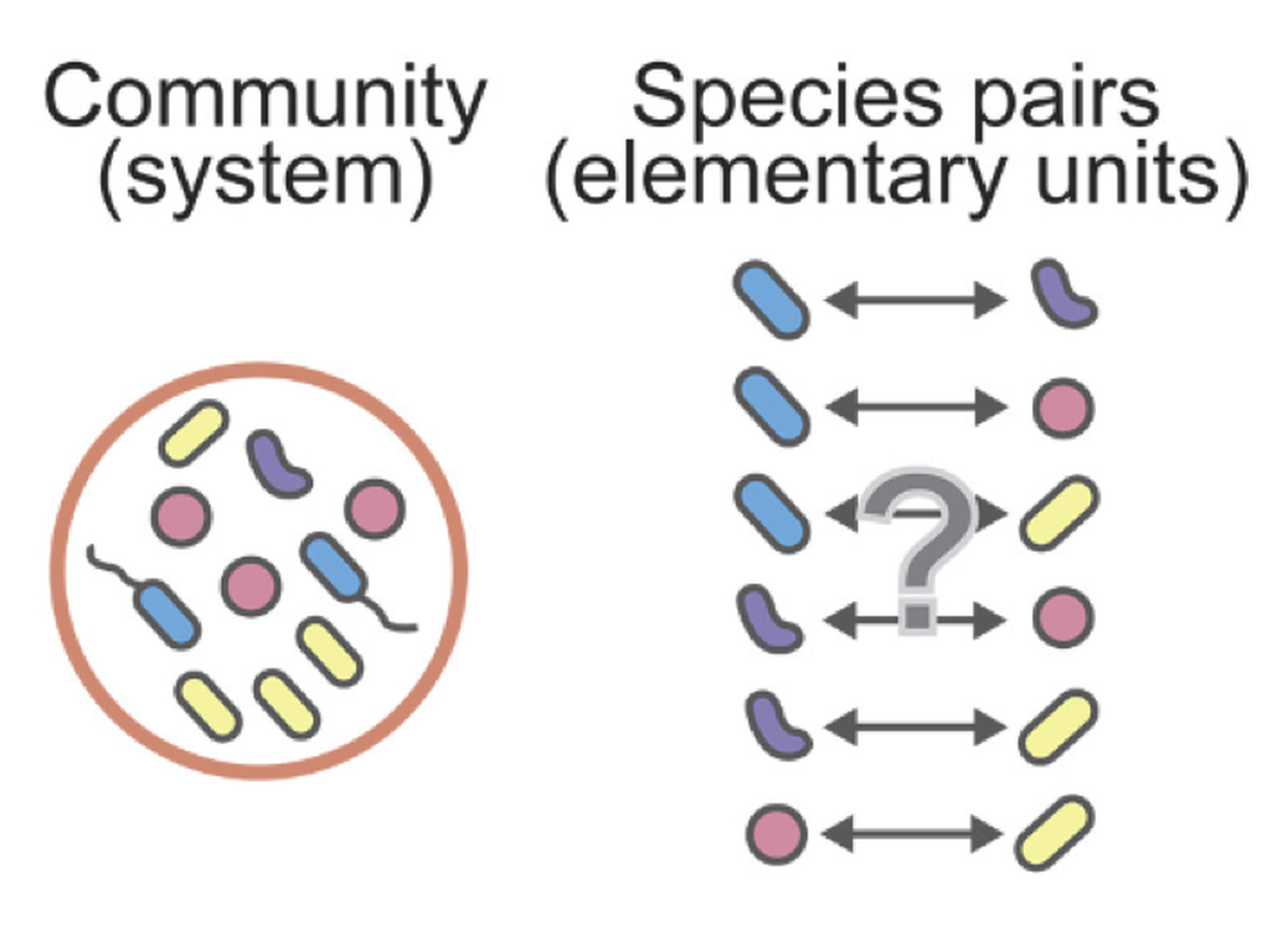
Why do diverse microbial consortia coexist, and can such coexistence be explained by simple pairwise principles?
Coexistence arises as an emergent property of microbial communities— systems that remain stable even when their constituent pairs cannot.
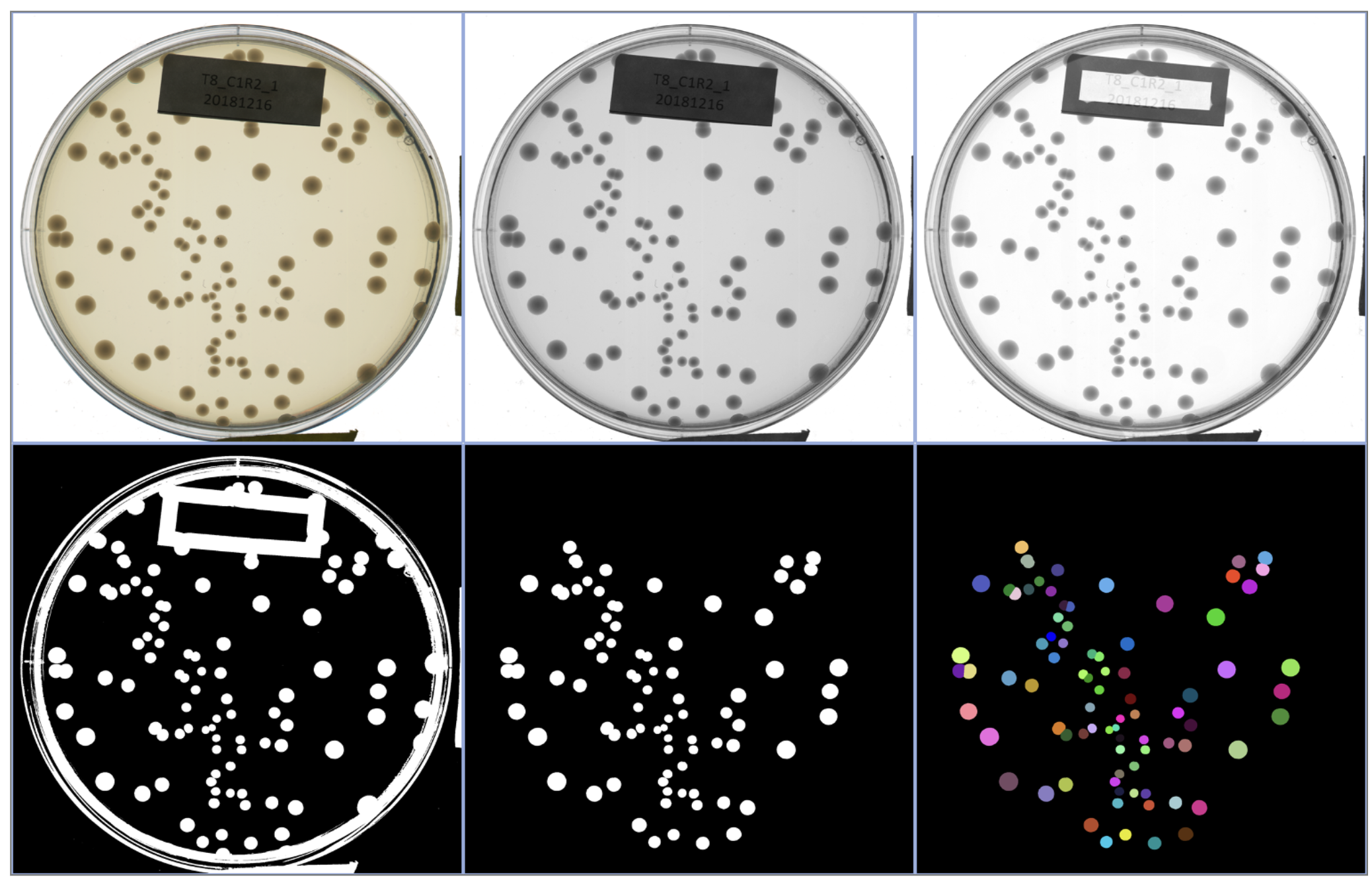
Synthetic communities, competitive assays, machine-learning–based image analysis.
Community ecology, microbial ecology, complex system.
We combine experiments and theory to explore how stability emerges from complex microbial interactions. Our results reveal that coexistence is not reducible to pairwise outcomes but is an emergent, collective feature of multispecies communities.
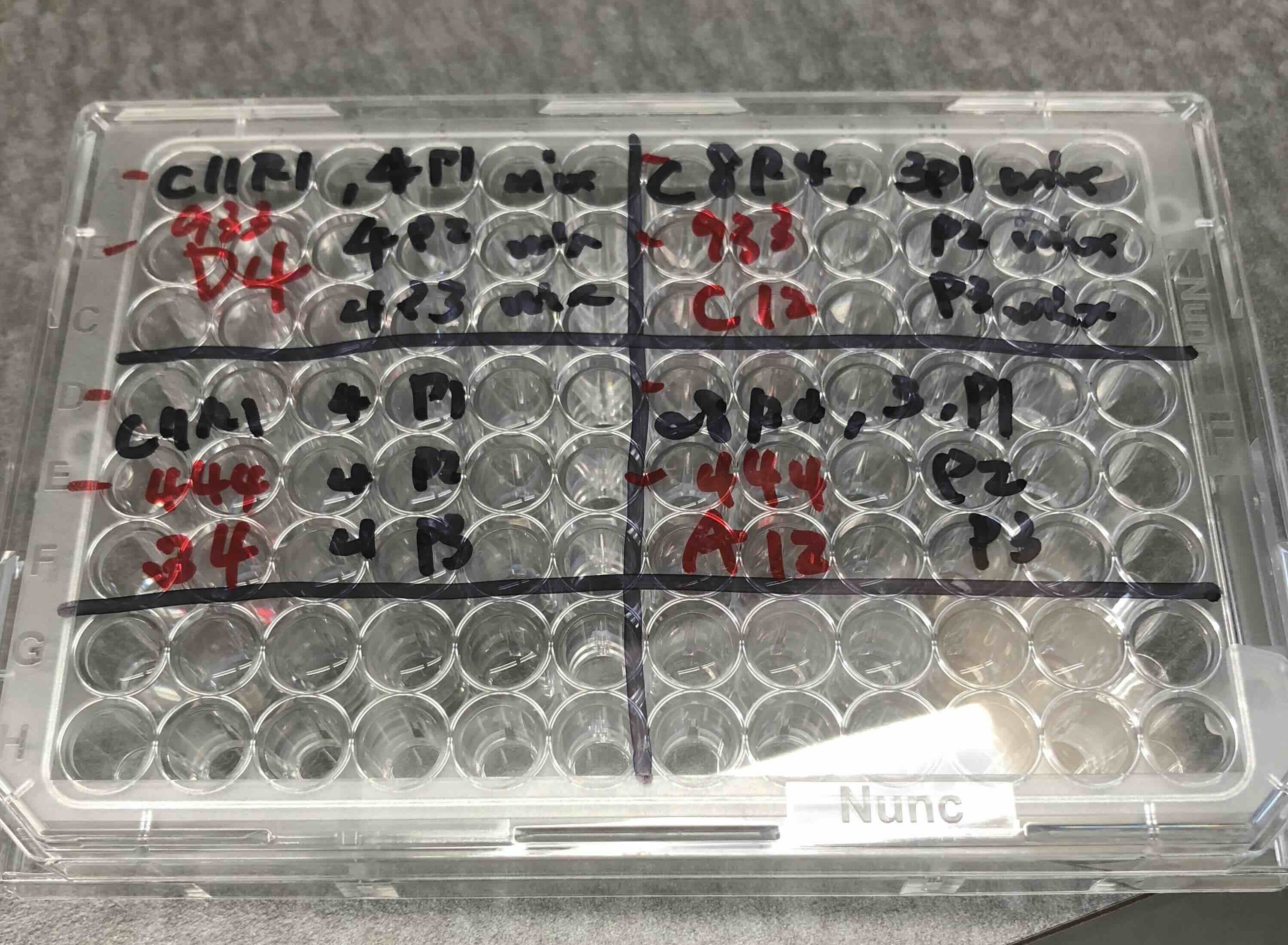
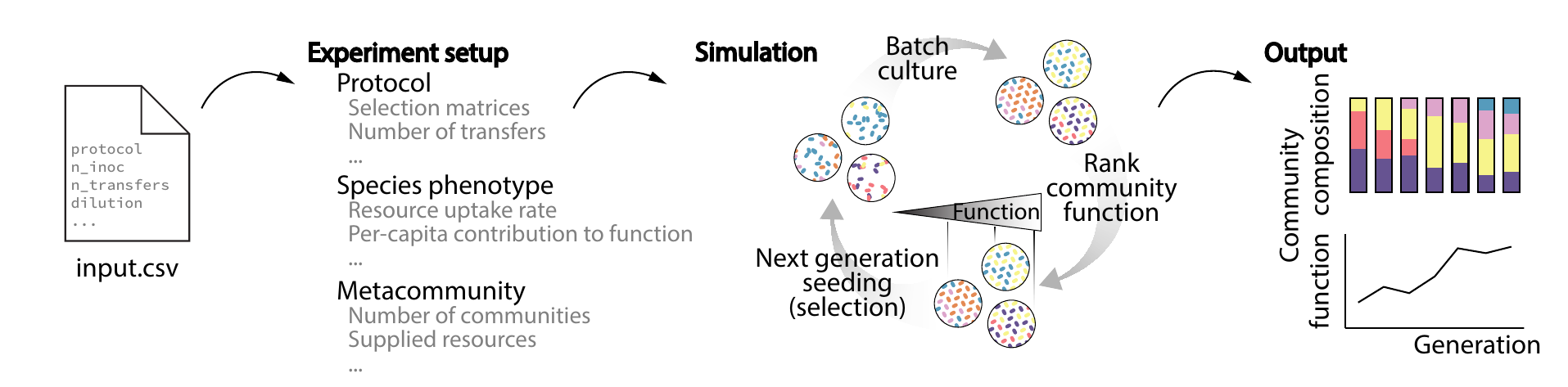
Microbiome Engineering & Artificial Selection
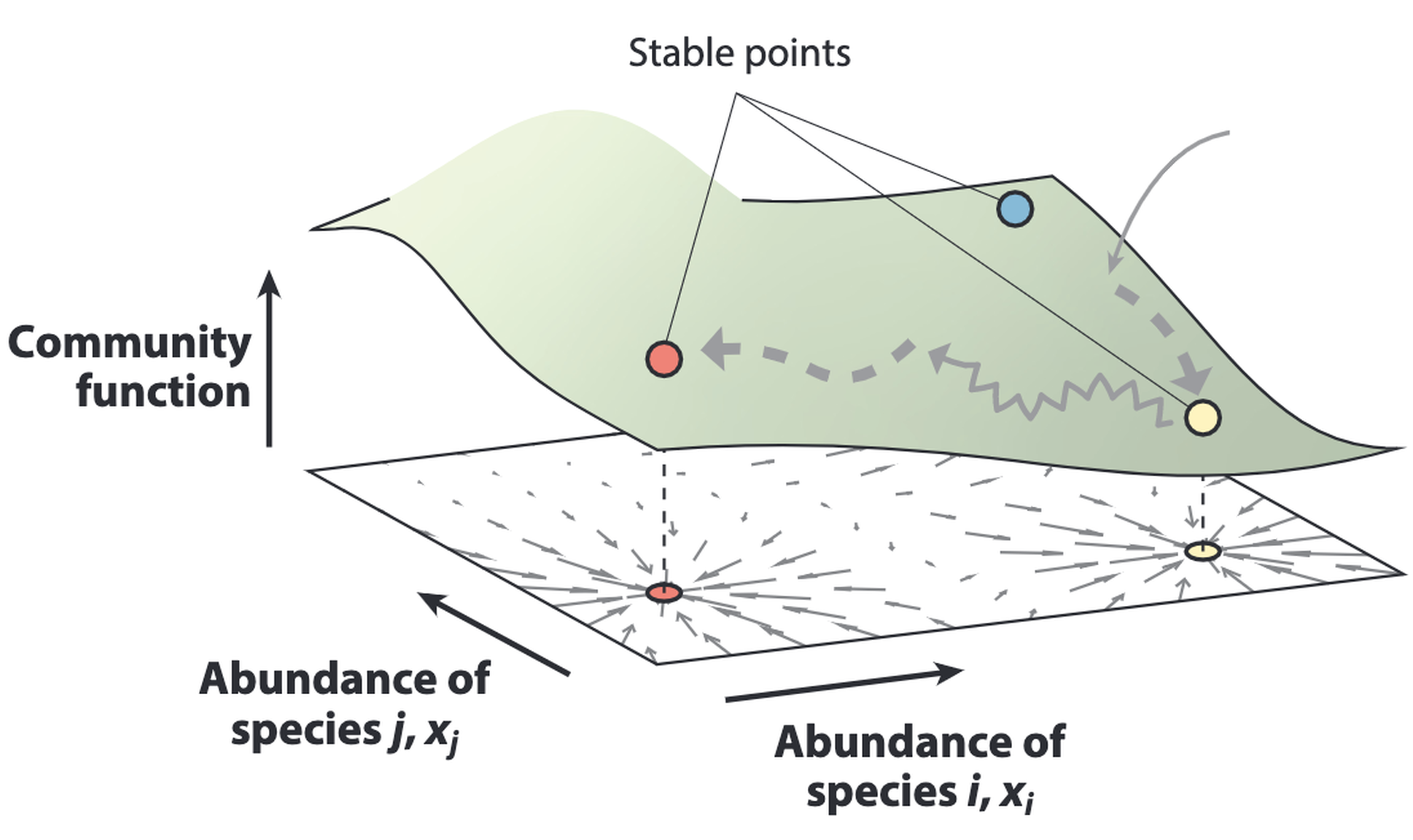
How can microbial communities be evolved or engineered as functional units through artificial selection?
Community-level functions can improve across selection cycles when heritable variation is maintained or replenished.
Experimental evolution, metabolic assays, computational modeling.
Microbiome engineering, artificial selection, directed evolution.
Chang et al. 2020 · Chang et al. 2021
We treat microbial consortia as evolving entities, subject to artificial selection on desired community traits. By combining experimental evolution and modeling, we develop frameworks for optimizing microbiome function and stability.
Studying Plants and Microbes Be Like: